Tools developed by chemists at NCI Frederick have at last made it possible to see how some of the most common molecules affect cells’ behavior in cancer and disease.
The sugar molecules, known as glycans, coat the outside of cells and play an important role in many biological functions. But they’re often altered in disease states, like cancer. This makes them a potentially useful target for medical research, diagnostics, and drug development.
Studying these molecules, because they can’t directly be sequenced, has been challenging. Unlike proteins, glycans are not genetically encoded, so common methods of molecular research either don’t work or aren’t specific enough for meaningful interpretation. New research from NCI Frederick’s Chemical Biology Laboratory (CBL) may make it easier.
What Is Glycomic-Seq?
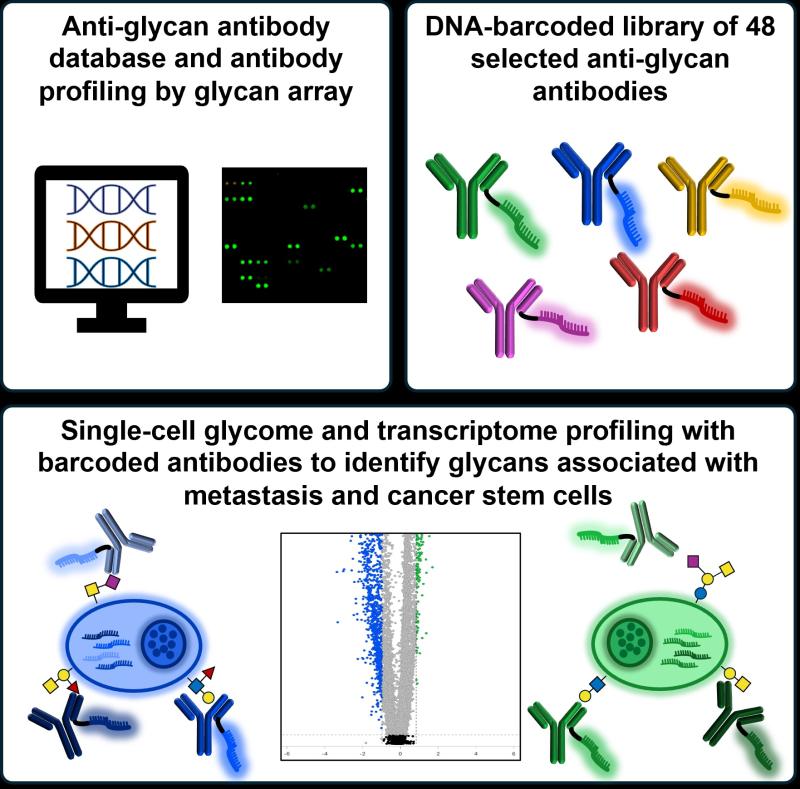
In a paper published in Cell Chemical Biology, these researchers describe their new tool, Glycomic-Seq, that enables many glycans to be profiled at once while simultaneously profiling the RNA transcripts produced by the cell, which tells them which genes are being transcribed in those same cells, all at the single-cell level.
To be able to identify glycans more easily, a team led by Jeffrey Gildersleeve, Ph.D., spent the months of remote work during the pandemic compiling a library of antibody sequences known to target specific glycans.
“Assembling the database was painstaking work that required each antibody sequence and associated metadata to be entered by hand, manually checked for accuracy, and curated,” said Gildersleeve, who is the head of the CBL’s Chemical Glycobiology Section.
The largest phase of the project came next, spearheaded by Samantha Marglous, Aneesa Bhakta, and Kara Gillman, a trio of postbaccalaureate fellows working for Gildersleeve.
How Was It Developed?
Marglous and Gillman generated and characterized a subset of about 150 antibodies from the database. Using this antibody library, Marglous created and attached DNA “barcodes” to the anti-glycan antibodies, which were then introduced to cells. If they saw the DNA barcode for a certain antibody when profiling a cell, they knew the cell had the corresponding glycan. Marglous designed and carried out the first single-cell profiling experiment.
Gildersleeve said figuring out how to make this process work was very difficult, but the resulting tool, Glycomic-Seq, integrated seamlessly with existing barcoding techniques and allowed glycan information and RNA transcripts to be collected together.
“It takes a lot of courage and resilience to tackle development of new technologies and forge a path forward, and the team deserve a ton of credit for overcoming numerous obstacles and figuring out how to successfully complete the project,” said Gildersleeve.
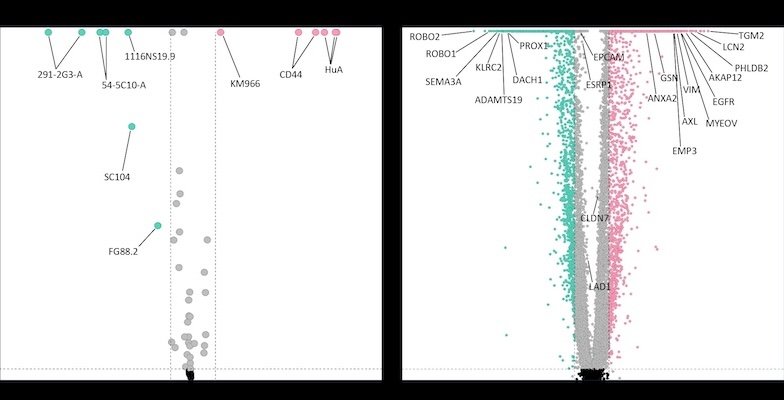
After the sequencing, Bhakta performed bioinformatics analyses on the results. This enabled the team to correlate glycan and RNA expression patterns to understand glycan expression on different cell types.
What Are the Next Steps?
In the end, the team used their technique to profile RNA and glycans on the surface of individual cells in two cancer cell lines, demonstrating the platform’s efficacy and revealing glycans associated with different cancer stem cell subsets.
Gildersleeve said the success of this platform opens “a wide range of new avenues of investigation.”
Researchers know little about glycan production in different cell types or how cell-surface glycans vary among cells. The team is interested in the potential to use their approach to study possible targets for cancer.
“Our hope is that Glycomic-Seq can be applied to understand how glycans are involved in cancer biology and to identify glycans associated with cancer that could be targeted therapeutically,” said Marglous.
Karolina Wilk is a technical editor in SPGM, where she writes for NCI Frederick and Frederick National Laboratory’s news outlets and edits scientific manuscripts, corporate documentation, and other writing. SPGM is the creative services department and hub for editing, illustration, graphic design, formatting, and multimedia training and support.