Nadya Tarasova, Ph.D., clicked her mouse a few times, repositioning some letters and lines on her computer screen. The program on display looked like it could be Photoshop’s distant cousin, but even an uninitiated onlooker could’ve seen this wasn’t artwork.
It was SLICE (short for “SMARTS and Logic in Chemistry”), one of the world’s first—and newest—chemistry coding programs to have a graphical interface.
With SLICE, scientists can design chemical models and run billions of virtual chemical reactions without having to write lengthy computer code, unlike with many previous programs that depend on the user having coding experience. Data also indicate SLICE completes these virtual syntheses much faster than at least one predecessor.
That means computer studies to model new potential drugs and their targets become faster and easier to perform, shortening the time needed to get compounds into laboratory studies and human clinical trials.
Stefi Nouleho Ilemo, Ph.D., former postdoctoral fellow in NCI Frederick’s Cancer Innovation Laboratory, and Victorien Delanée, Ph.D., former postdoctoral fellow in the Chemical Biology Laboratory, coded SLICE to advance, empower, and unburden computational chemistry, the field of using computers to study chemicals and their interactions.
“Two very talented postdocs wrote the code for the program, although the effort included many other people,” said Tarasova, senior associate scientist in the Cancer Innovation Laboratory.
SLICE was reported in the Journal of Chemoinformatics and is available for free via GitHub.
No Code-Writing Required
Through SLICE, any user with sufficient chemistry expertise can enter data for chemicals and their products and synthesize vast databases, Tarasova said.
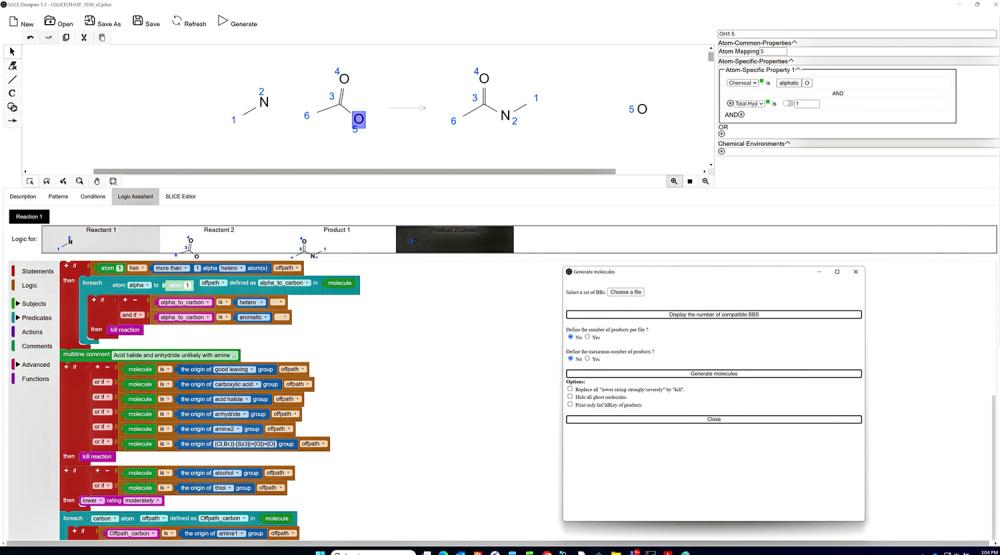
Users sketch chemical reactions through SLICE’s graphical interface. Menus give options to define the properties of every atom. The SLICE Logic Assistant—modular, movable instructions resembling puzzle pieces—lets users dictate further parameters the molecules must satisfy.
These features make it possible to tailor reactions and avoid junk results, without painstaking manual input through CHMTRN or a similar programming language. Programming a single type of synthetic chemistry in traditional programs can require 10–12 pages of code because the user must write out the properties of the chemicals involved.
Yet with SLICE, “we can easily program novel chemistry, and novel chemistry means novel classes of compounds, novel scaffolds, better diversity,” Tarasova said.
Better diversity implies a higher probability of finding effective drugs, she said.
SLICE as a Shortcut
SLICE’s speed also improves on existing programs. It vastly outperformed its predecessor, SAVI, in an NCI Frederick test synthesis using four popular synthetic methods applied to thousands of chemicals, with millions of resulting compounds per set.
Scientists ran SLICE and SAVI in an equal way: on the same computer system with the same specifications, using the same computational resources on the National Institutes of Health Biowulf supercomputer. At its slowest, SLICE was nearly 13 times quicker. At its fastest, it was almost 200 times faster, finishing in two hours what took SAVI 11 days to complete.
“If your process runs a hundred times faster, that’s a hundred times less used resources,” Tarasova said. “So, you are making those nodes that you are sitting on on Biowulf available to somebody else. … It allows you to adjust your research very fast, too.”
Speed lets scientists execute both more and larger analyses. That helps them partition the gargantuan realm of potential drugs into explorable chunks.
And that field is unfathomably massive. There are 1063 drug-like compounds predicted to exist. That’s 10 followed by 63 zeroes, a sum so huge that if the 10 nonillion living organisms on Earth were added to the 200 billion trillion stars in the universe, the total still wouldn’t even come close to comparing.
Any one of these compounds might become a new medicine, but scientists have so far studied only 130 million of them, “less than a drop in the ocean,” Tarasova said.
The difficulty of performing large-scale analyses, whether due to limited resources, time, or coding expertise, has contributed to that. SLICE doesn’t fix it all, but by allowing chemists to access the most relevant parts of the chemical space faster and easier, it aims to help.
“It is a shortcut that can significantly accelerate virtual drug discovery,” Tarasova said.
The Cancer Innovation Laboratory, the Chemical Biology Laboratory, Frederick National Laboratory for Cancer Research, and Philip Judson (LHASA Ltd, England) contributed to the study. SLICE is available for one-click installation on Windows, Mac, and Linux via GitHub. Users with questions can contact Tarasova.
Samuel Lopez leads the editorial team in Scientific Publications, Graphics & Media (SPGM). He writes for newsletters; informally serves as an institutional historian; and edits scientific manuscripts, corporate documents, and sundry other written media. SPGM is the creative services department and hub for editing, illustration, graphic design, formatting, and multimedia.