The old adage that says two heads are better than one certainly seems true for Mitchell Ho, Ph.D., a senior investigator in the Center for Cancer Research (CCR), and Xiaolin Wu, Ph.D., a principal scientist in the Genomics Technology Laboratory, a CCR Core at the Frederick National Laboratory.
Together, these two scientists are using next-generation genomics technology to develop, in an animal model, a chimeric antigen receptor (CAR)-T cell therapy that might help patients with hepatocellular carcinoma (HCC), the most common type of liver cancer.
Their study, published in Gastroenterology, shows that injections of CAR (hYP7) T cells eliminated tumors in 66% of samples in an animal model of human disease within three weeks and continued to do so even after a re-challenge with tumor cells—making this a potentially promising therapeutic for HCC.
A Molecular Approach: “The Devil Is in the Details”
While CAR-T cell therapy has been used to treat blood cancers like leukemia, it has not been successful in treating solid tumors, which are more difficult for CAR T cells to penetrate. One major challenge has been the lack of a molecular target.
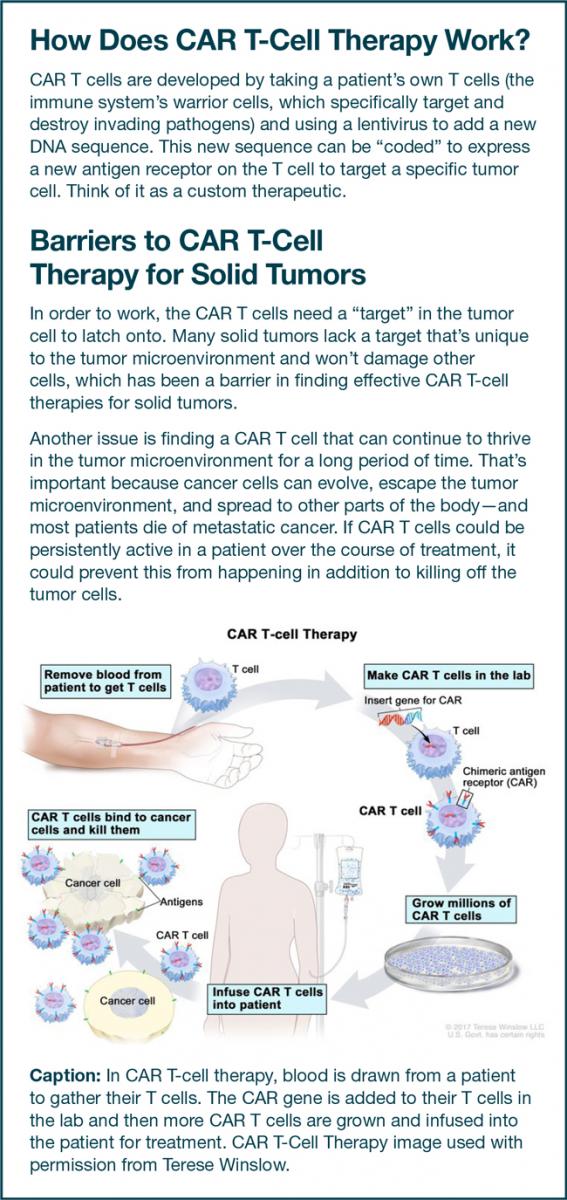 But Ho and his team think they are on to something. They’re attempting to target GPC3, a protein that appears only in embryos or tumors but not normal adult tissue and is highly expressed in HCC, making it a reliable target. Ho identified two potential antibodies, YP7 and HN3, that might home in on GPC3 and sought to test their effectiveness.
But Ho and his team think they are on to something. They’re attempting to target GPC3, a protein that appears only in embryos or tumors but not normal adult tissue and is highly expressed in HCC, making it a reliable target. Ho identified two potential antibodies, YP7 and HN3, that might home in on GPC3 and sought to test their effectiveness.
He and his team first took cells from healthy donors and patients with HCC to create CAR T cells based on humanized versions of the two potential antibodies (hYP7 and HN3). The two types of CAR T cells, as well as a control T cell, were tested as treatments against HCC in human tumor xenograft models.
The researchers needed to analyze tissue samples on a molecular level to see if the CAR T cells were persistent in the tumor microenvironment and determine which therapy was effective. “The devil is always in the details [when it comes to biology]. If we don’t know gene sequences in cells or tissues, as a molecular biologist, I just don’t feel confident that we understand the mechanism,” said Ho.
Joining Forces: More Expertise is Better
In the study’s original design, Ho’s team was planning to use a cumbersome method—tissue staining—to quantify the CAR T cells remaining in their samples. With that method, tissue samples have to be painstakingly scrutinized one by one under the microscope to identify and quantify the cells.
The project was already under way when some friendly dialogue led to an unexpected improvement in the methodology. Ho and Wu, who have known each other for several years, were meeting about an unrelated project when their conversation casually moved to Ho’s CAR T-cell therapy project. Wu mentioned that, for a different CAR-T cell therapy study, he was using next-generation genomic technology. He explained that his method can quantify what Ho’s staining protocols were attempting to measure using only genomic DNA (not whole tissue samples) and can analyze the CAR T cells more rapidly and precisely.
After talking through the methodology with Wu, Ho’s team stopped the staining protocols and instead moved forward with Wu’s suggested technologies, digital droplet polymerase chain reaction (ddPCR) and vector integration site analysis (VISA).
“Each of the core facilities and those labs have expertise, and that’s what we need to make cancer biology research cutting-edge,” said Ho.
The benefits of this collaborative setup work both ways. Wu says working with researchers on the clinical side not only shows the potential of new applications for their work but also reveals which improvements in the technology would be helpful.
“I think, once you’re out of school, you have to be able to learn whatever you’re working on. … [Collaboration] is a great opportunity for us to learn new areas,” said Wu. It’s one of the benefits of the Genomics Technology Laboratory’s design as a core facility to support CCR scientists.
Next-Generation Genomics Technology
Polymerase chain reaction (PCR) takes a small DNA sample and amplifies it by millions to make it easier to study in detail. ddPCR is the latest improvement on this technology. In this process, a sample is separated into droplets, and PCR amplification occurs in each droplet. Because the sample is partitioned before amplification, scientists can reliably measure minute differences in the DNA sample. The process results in a precise measurement from a small sample—and it’s much faster. Wu’s team is able to process up to 96 samples a day.
For this project, they used ddPCR to determine what percentage of the cells in the sample were tumor cells versus CAR T cells after treatment. This could tell them in just a few days how effective the therapy was.
Once they determined that the CAR T-cell therapy was working, the team used VISA to figure out which of the CAR T cells in the experiment performed better. When CAR T cells are developed, the lentiviral vector’s integration into the T-cell genome is random. Since it’s unlikely that different CAR T cells would have the same integration site, each one acts as a sort of “barcode” that can identify a unique CAR T cell.
Wu began working with this kind of technology 15 years ago. Then, it was a great accomplishment to find 900 integration sites over a couple years. Using the current, improved version, they can easily map millions of integration sites in a short period of time.
The Future of CAR T-Cell Therapy for HCC
For Ho’s project, the technology allowed them to see that the CAR (hYP7) T cells remained in the tumor microenvironment for up to seven weeks. Plus, they were able to pinpoint the cells with enough precision to know the CAR (hYP7) T cells were effective even with different integration sites and with samples from different time periods—meaning that they could more confidently say this particular type of CAR T cell has the potential to be an effective therapy.
Ho hopes that this study is one step in the process toward more potent CAR T therapies. He is looking forward to the next step—the study is now moving forward with clinical trials at the NIH Clinical Center—and he’s interested to see whether the CAR T-cell therapy persists in patients throughout the course of the treatment.
He also notes that, besides efficacy, the safety of the treatment will need to be tested. It could still be a long time before this therapeutic is available to patients. In the meantime, Ho and his team were excited to find new technologies that could aid their experiment, and they are now applying these methods to other studies, as well.
“Technology is always moving [forward], and we’re always trying to look for new technologies to help us understand more precisely tumor biology and treatment,” said Ho.
For Wu and his team, the collaboration brings a welcome application to their data-driven work. “I just feel so excited every day to work on these different projects because I come from a background of doing basic research. … Every project [for NCI CCR] is exciting because we know it can benefit … patients,” said Wu.